обновлено 26.06.2022
R1b-PF7562 - субклад гаплогруппы R1b, исходящий из узла M269 и параллельный узлу L23.
Ареал R1b-PF7562
Частоты R1b-PF7562 (в источниках M269(xL23)):
Macedonia - 4/79 (5,06 %)
Albania - 11/223 (4,93 %)
Serbia - 7/235 (2,98 %)
Armenia - 5/176 (2,84 %)
Cyprus - 16/574 (2,79 %)
Laz - 1/36 (2,78 %)
Lezgins - 1/41 (2,44 %)
Italy - 26/1094 (2,38 %)
Tabasarans - 1/43 (2,33 %)
Greece - 8/347 (2,31 %)
Turkey - 25/1176 (2,13 %)
Crete - 4/193 (2,07 %)
Algeria - 2/102 (1,96 %)
Romania - 10/527 (1,9 %)
Bashkirs - 10/586 (1,71 %)
Herzegovina - 2/141 (1,42 %)
Bosnia - 1/78 (1,28 %)
Syria - 1/81 (1,23 %)
Sardinia - 28/2404 (1,16 %)
Switzerland - 2/175 (1,14 %)
Hungary - 4/370 (1,08 %)
Bulgaria - 10/931 (1,07 %)
Tunisia - 1/120 (0,83 %)
Slovenia - 4/501 (0,8 %)
Iran - 10/1303 (0,77 %)
Mongolia - 1/160 (0,63 %)
Palestinians - 1/170 (0,59 %)
Germany - 4/727 (0,55 %)
Portugal - 1/190 (0,53 %)
Denmark - 1/215 (0,47 %)
Ukraine - 2/596 (0,34 %)
Georgia - 1/345 (0,29 %)
Jordan - 1/392 (0,26 %)
Russians - 3/1225 (0,24 %)
Afghanistan - 1/507 (0,2 %)
France - 1/535 (0,19 %)
Netherlands - 3/2172 (0,14 %)
Poland - 1/772 (0,13 %)
Croatia - 1/828 (0,12 %)
Spain - 1/1122 (0,09 %)
Источники:
Crete - 4/193 (2,07 %)
Algeria - 2/102 (1,96 %)
Romania - 10/527 (1,9 %)
Bashkirs - 10/586 (1,71 %)
Herzegovina - 2/141 (1,42 %)
Bosnia - 1/78 (1,28 %)
Syria - 1/81 (1,23 %)
Sardinia - 28/2404 (1,16 %)
Switzerland - 2/175 (1,14 %)
Hungary - 4/370 (1,08 %)
Bulgaria - 10/931 (1,07 %)
Tunisia - 1/120 (0,83 %)
Slovenia - 4/501 (0,8 %)
Iran - 10/1303 (0,77 %)
Mongolia - 1/160 (0,63 %)
Palestinians - 1/170 (0,59 %)
Germany - 4/727 (0,55 %)
Portugal - 1/190 (0,53 %)
Denmark - 1/215 (0,47 %)
Ukraine - 2/596 (0,34 %)
Georgia - 1/345 (0,29 %)
Jordan - 1/392 (0,26 %)
Russians - 3/1225 (0,24 %)
Afghanistan - 1/507 (0,2 %)
France - 1/535 (0,19 %)
Netherlands - 3/2172 (0,14 %)
Poland - 1/772 (0,13 %)
Croatia - 1/828 (0,12 %)
Spain - 1/1122 (0,09 %)
Источники:
The genetic structure of the Turkish population reveals high levels of variation and admixture, Kars 2021
Y-chromosome and Surname Analyses for Reconstructing Past Population Structures: The Sardinian Population as a Test Case, Grugni 2019
The Dutch Y-chromosomal landscape, Altena 2019
Reconstructing the genetic history of Italians: new insights from a male (Y-chromosome) perspective, Grugni 2018
Genetic characterization of Balkars and Karachays according to the variability of the Y chromosome, Dzhaubermezov 2017
Is there a Finno-Ugric component in the gene pool of Russians from Yaroslavl oblast? Evidence from Y-chromosome, Chukhryaeva 2017
Genetic differentiation between upland and lowland populations shapes the Y-chromosomal landscape of West Asia, Balanovsky 2017
Genetic heritage of Croatians in the Southeastern European gene pool—Y chromosome analysis of the Croatian continental and Island population, Šarac 2016
Y chromosome diversity in a linguistic isolate (Mirandese, NE Portugal), Marques 2016
Y-chromosome phylogeographic analysis of the Greek-Cypriot population reveals elements consistent with Neolithic and Bronze Age settlements, Voskarides 2016
Coevolution of genes and languages and high levels of population structure among the highland populations of Daghestan, Karafet 2015
Origins, admixture and founder lineages in European Roma, Martínez-Cruz 2015
Shared language, diverging genetic histories: high-resolution analysis of Y-chromosome variability in Calabrian and Sicilian Arbereshe, Sarno 2015
Large-scale recent expansion of European patrilineages shown by population resequencing, Batini 2015
Detection of phylogenetically informative polymorphisms in the entire euchromatic portion of human Y chromosome from a Sardinian sample, Francalacci 2015
Mitochondrial DNA and Y-chromosome structure at the mediterranean and atlantic façades of the iberian peninsula, Santos 2013
Afghan Hindu Kush: Where Eurasian Sub-Continent Gene Flows Converge, Di Cristofaro 2013
Low-Pass DNA Sequencing of 1200 Sardinians Reconstructs European Y-Chromosome Phylogeny, Francalacci 2013
The paternal perspective of the Slovenian population and its relationship with other populations, Zupan 2013
Y-Chromosome Diversity in Modern Bulgarians: New Clues about Their Ancestry, Karachanak 2013
Introducing the Algerian Mitochondrial DNA and Y-Chromosome Profiles into the North African Landscape, Bekada 2013
Contemporary paternal genetic landscape of Polish and German populations: from early medieval Slavic expansion to post-World War II resettlements, Rębała 2012
Pasture Names with Romance and Slavic Roots Facilitate Dissection of Y Chromosome Variation in an Exclusively German-Speaking Alpine Region, Niederstätter 2012
Ancient Migratory Events in the Middle East: New Clues from the Y-Chromosome Variation of Modern Iranians, Grugni 2012
High levels of Paleolithic Y-chromosome lineages characterize Serbia, Regueiro 2012
Neolithic patrilineal signals indicate that the Armenian plateau was repopulated by agriculturalists, Herrera 2011
A major Y-chromosome haplogroup R1b Holocene era founder effect in Central and Western Europe, Myres 2010
Распространение R1b-PF7562 совпадает в основном с распространением субклада R1b-Z2103.
Ареал R1b-Z2103
Y-chromosome and Surname Analyses for Reconstructing Past Population Structures: The Sardinian Population as a Test Case, Grugni 2019
The Dutch Y-chromosomal landscape, Altena 2019
Reconstructing the genetic history of Italians: new insights from a male (Y-chromosome) perspective, Grugni 2018
Genetic characterization of Balkars and Karachays according to the variability of the Y chromosome, Dzhaubermezov 2017
Is there a Finno-Ugric component in the gene pool of Russians from Yaroslavl oblast? Evidence from Y-chromosome, Chukhryaeva 2017
Genetic differentiation between upland and lowland populations shapes the Y-chromosomal landscape of West Asia, Balanovsky 2017
Genetic heritage of Croatians in the Southeastern European gene pool—Y chromosome analysis of the Croatian continental and Island population, Šarac 2016
Y chromosome diversity in a linguistic isolate (Mirandese, NE Portugal), Marques 2016
Y-chromosome phylogeographic analysis of the Greek-Cypriot population reveals elements consistent with Neolithic and Bronze Age settlements, Voskarides 2016
Coevolution of genes and languages and high levels of population structure among the highland populations of Daghestan, Karafet 2015
Origins, admixture and founder lineages in European Roma, Martínez-Cruz 2015
Shared language, diverging genetic histories: high-resolution analysis of Y-chromosome variability in Calabrian and Sicilian Arbereshe, Sarno 2015
Large-scale recent expansion of European patrilineages shown by population resequencing, Batini 2015
Detection of phylogenetically informative polymorphisms in the entire euchromatic portion of human Y chromosome from a Sardinian sample, Francalacci 2015
Mitochondrial DNA and Y-chromosome structure at the mediterranean and atlantic façades of the iberian peninsula, Santos 2013
Afghan Hindu Kush: Where Eurasian Sub-Continent Gene Flows Converge, Di Cristofaro 2013
Low-Pass DNA Sequencing of 1200 Sardinians Reconstructs European Y-Chromosome Phylogeny, Francalacci 2013
The paternal perspective of the Slovenian population and its relationship with other populations, Zupan 2013
Y-Chromosome Diversity in Modern Bulgarians: New Clues about Their Ancestry, Karachanak 2013
Introducing the Algerian Mitochondrial DNA and Y-Chromosome Profiles into the North African Landscape, Bekada 2013
Contemporary paternal genetic landscape of Polish and German populations: from early medieval Slavic expansion to post-World War II resettlements, Rębała 2012
Pasture Names with Romance and Slavic Roots Facilitate Dissection of Y Chromosome Variation in an Exclusively German-Speaking Alpine Region, Niederstätter 2012
Ancient Migratory Events in the Middle East: New Clues from the Y-Chromosome Variation of Modern Iranians, Grugni 2012
High levels of Paleolithic Y-chromosome lineages characterize Serbia, Regueiro 2012
Neolithic patrilineal signals indicate that the Armenian plateau was repopulated by agriculturalists, Herrera 2011
A major Y-chromosome haplogroup R1b Holocene era founder effect in Central and Western Europe, Myres 2010
Распространение R1b-PF7562 совпадает в основном с распространением субклада R1b-Z2103.
Ареал R1b-Z2103
Частоты R1b-Z2103 (в тех же источниках L23(xL51)):
Armenia - 45/176 (25,57 %)
Bashkirs - 126/586 (21,5 %)
Dagestan - 87/724 (12,02 %)
Komis (Perm Oblast) - 7/61 (11,48 %)
Kosovo - 13/114 (11,4 %)
Turkey - 129/1176 (10,97 %)
Albania - 22/223 (9,87 %)
Iran - 106/1303 (8,14 %)
Iraq - 2/28 (7,14 %)
Greece - 21/347 (6,05 %)
Cyprus - 34/574 (5,92 %)
Czechia - 5/87 (5,75 %)
Switzerland - 10/175 (5,71 %)
Bulgaria - 50/931 (5,37 %)
Romania - 28/527 (5,31 %)
Syria - 4/81 (4,94 %)
Hungary - 16/370 (4,32 %)
Sweden - 6/139 (4,32 %)
Italy - 46/1094 (4,2 %)
Tatars - 5/119 (4,2 %)
Crete - 8/193 (4,15 %)
Ossetians - 6/157 (3,82 %)
Udmurts - 2/54 (3,7 %)
Slovakia - 22/601 (3,66 %)
Croatia - 30/828 (3,62 %)
Slovenia - 18/501 (3,59 %)
Belarus - 7/205 (3,41 %)
Abazins - 3/89 (3,37 %)
Saudi Arabia - 1/32 (3,13 %)
Uyghurs - 2/66 (3,03 %)
Afghanistan - 15/507 (2,96 %)
Russians - 36/1225 (2,94 %)
Kabardians - 4/141 (2,84 %)
Poland - 21/772 (2,72 %)
Uzbekistan - 2/74 (2,7 %)
Kyrgyzstan - 4/150 (2,67 %)
Georgia - 9/345 (2,61 %)
Pakistan - 12/461 (2,6 %)
Karachays - 5/195 (2,56 %)
Turkmenistan - 1/46 (2,17 %)
Serbia - 5/235 (2,13 %)
Portugal - 4/190 (2,11 %)
Ukraine - 12/596 (2,01 %)
Balkars - 7/371 (1,89 %)
Denmark - 4/215 (1,86 %)
Austria - 5/288 (1,74 %)
Chuvashes - 2/117 (1,71 %)
France - 9/535 (1,68 %)
Lusatians - 2/123 (1,63 %)
Germany - 11/727 (1,51 %)
Herzegovina - 2/141 (1,42 %)
Britain - 2/153 (1,31 %)
Palestinians - 2/170 (1,18 %)
Netherlands - 24/2172 (1,1 %)
Jordan - 4/392 (1,02 %)
India - 2/214 (0,93 %)
Sardinia - 22/2404 (0,92 %)
Ireland - 1/119 (0,84 %)
Spain - 9/1122 (0,8 %)
Cherkess - 1/126 (0,79 %)
Mongolia - 1/160 (0,63 %)
Karelians - 1/194 (0,52 %)
Estonia - 1/210 (0,48 %)
Схожесть ареалов PF7562 и Z2103 указывает на то, что в далёком прошлом эти ветви, очевидно, составляли одну общность. В связи с этим интересно посмотреть на суммарный ареал обоих субкладов, то есть ареал R1b-M269(xL51).
Ареал R1b-M269(xL51)
Albania - 22/223 (9,87 %)
Iran - 106/1303 (8,14 %)
Iraq - 2/28 (7,14 %)
Greece - 21/347 (6,05 %)
Cyprus - 34/574 (5,92 %)
Czechia - 5/87 (5,75 %)
Switzerland - 10/175 (5,71 %)
Bulgaria - 50/931 (5,37 %)
Romania - 28/527 (5,31 %)
Syria - 4/81 (4,94 %)
Hungary - 16/370 (4,32 %)
Sweden - 6/139 (4,32 %)
Italy - 46/1094 (4,2 %)
Tatars - 5/119 (4,2 %)
Crete - 8/193 (4,15 %)
Ossetians - 6/157 (3,82 %)
Udmurts - 2/54 (3,7 %)
Slovakia - 22/601 (3,66 %)
Croatia - 30/828 (3,62 %)
Slovenia - 18/501 (3,59 %)
Belarus - 7/205 (3,41 %)
Abazins - 3/89 (3,37 %)
Saudi Arabia - 1/32 (3,13 %)
Uyghurs - 2/66 (3,03 %)
Afghanistan - 15/507 (2,96 %)
Russians - 36/1225 (2,94 %)
Kabardians - 4/141 (2,84 %)
Poland - 21/772 (2,72 %)
Uzbekistan - 2/74 (2,7 %)
Kyrgyzstan - 4/150 (2,67 %)
Georgia - 9/345 (2,61 %)
Pakistan - 12/461 (2,6 %)
Karachays - 5/195 (2,56 %)
Turkmenistan - 1/46 (2,17 %)
Serbia - 5/235 (2,13 %)
Portugal - 4/190 (2,11 %)
Ukraine - 12/596 (2,01 %)
Balkars - 7/371 (1,89 %)
Denmark - 4/215 (1,86 %)
Austria - 5/288 (1,74 %)
Chuvashes - 2/117 (1,71 %)
France - 9/535 (1,68 %)
Lusatians - 2/123 (1,63 %)
Germany - 11/727 (1,51 %)
Herzegovina - 2/141 (1,42 %)
Britain - 2/153 (1,31 %)
Palestinians - 2/170 (1,18 %)
Netherlands - 24/2172 (1,1 %)
Jordan - 4/392 (1,02 %)
India - 2/214 (0,93 %)
Sardinia - 22/2404 (0,92 %)
Ireland - 1/119 (0,84 %)
Spain - 9/1122 (0,8 %)
Cherkess - 1/126 (0,79 %)
Mongolia - 1/160 (0,63 %)
Karelians - 1/194 (0,52 %)
Estonia - 1/210 (0,48 %)
Схожесть ареалов PF7562 и Z2103 указывает на то, что в далёком прошлом эти ветви, очевидно, составляли одну общность. В связи с этим интересно посмотреть на суммарный ареал обоих субкладов, то есть ареал R1b-M269(xL51).
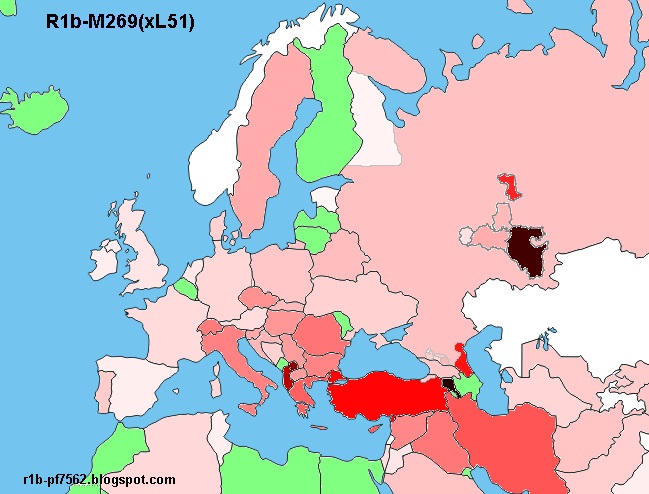
Ареал R1b-M269(xL51)
Частоты R1b-M269(xL51):
Armenia - 50/176 (28,41 %)
Bashkirs - 136/586 (23,21 %)
Kosovo - 22/114 (19,3 %)
Albania - 33/223 (14,8 %)
Turkey - 154/1176 (13,1 %)
Dagestan - 89/724 (12,29 %)
Komis (Perm Oblast) - 7/61 (11,48 %)
Iran - 116/1303 (8,9 %)
Cyprus - 50/574 (8,71 %)
Greece - 29/347 (8,36 %)
Romania - 38/527 (7,21 %)
Iraq - 2/28 (7,14 %)
Switzerland - 12/175 (6,86 %)
Italy - 72/1094 (6,58 %)
Bulgaria - 60/931 (6,44 %)
Crete - 12/193 (6,22 %)
Syria - 5/81 (6,17 %)
Czechia - 5/87 (5,75 %)
Hungary - 20/370 (5,41 %)
Serbia - 12/235 (5,11 %)
Macedonia - 4/79 (5,06 %)
Slovenia - 22/501 (4,39 %)
Sweden - 6/139 (4,32 %)
Tatars - 5/119 (4,2 %)
Ossetians - 6/157 (3,82 %)
Croatia - 31/828 (3,74 %)
Udmurts - 2/54 (3,7 %)
Slovakia - 22/601 (3,66 %)
Belarus - 7/205 (3,41 %)
Abazins - 3/89 (3,37 %)
Russians - 39/1225 (3,18 %)
Afghanistan - 16/507 (3,16 %)
Saudi Arabia - 1/32 (3,13 %)
Uyghurs - 2/66 (3,03 %)
Georgia - 10/345 (2,9 %)
Poland - 22/772 (2,85 %)
Herzegovina - 4/141 (2,84 %)
Kabardians - 4/141 (2,84 %)
Laz - 1/36 (2,78 %)
Uzbekistan - 2/74 (2,7 %)
Kyrgyzstan - 4/150 (2,67 %)
Portugal - 5/190 (2,63 %)
Pakistan - 12/461 (2,6 %)
Karachays - 5/195 (2,56 %)
Ukraine - 14/596 (2,35 %)
Denmark - 5/215 (2,33 %)
Turkmenistan - 1/46 (2,17 %)
Sardinia - 50/2404 (2,08 %)
Germany - 15/727 (2,06 %)
Algeria - 2/102 (1,96 %)
Balkars - 7/371 (1,89 %)
France - 10/535 (1,87 %)
Palestinians - 3/170 (1,76 %)
Austria - 5/288 (1,74 %)
Chuvashes - 2/117 (1,71 %)
Lusatians - 2/123 (1,63 %)
Britain - 2/153 (1,31 %)
Jordan - 5/392 (1,28 %)
Bosnia - 1/78 (1,28 %)
Mongolia - 2/160 (1,25 %)
Netherlands - 27/2172 (1,24 %)
India - 2/214 (0,93 %)
Spain - 10/1122 (0,89 %)
Ireland - 1/119 (0,84 %)
Tunisia - 1/120 (0,83 %)
Cherkess - 1/126 (0,79 %)
Karelians - 1/194 (0,52 %)
Estonia - 1/210 (0,48 %)
https://discover.familytreedna.com/y-dna/R-PF7562/tree
https://www.familytreedna.com/public/y-dna-haplotree/R;name=R-PF7562
https://www.yfull.com/tree/R-PF7562/




-1.jpg)